| dc.contributor.author | Lohr, Chaelynne | |
| dc.contributor.author | Fu, Jun | |
| dc.contributor.author | Allen, Robert | |
| dc.date.accessioned | 2023-09-12T16:44:58Z | |
| dc.date.available | 2023-09-12T16:44:58Z | |
| dc.date.issued | 2022-02-18 | |
| dc.identifier | ouhd_Lohr_detectionofRNAmethylation_2022 | |
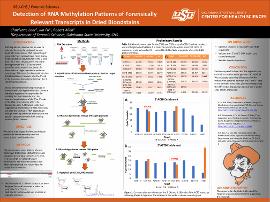
| dc.identifier.citation | Lohr, C., Fu, J., & Allen, R. (2022, February 18). Detection of RNA methylation patterns of forensically relevant transcripts in dried bloodstains. Poster presented at Research Days at Oklahoma State University Center for Health Sciences, Tulsa, Ok | |
| dc.identifier.uri | https://hdl.handle.net/11244/339528 | |
| dc.description.abstract | RNA degradation kinetics can be used to estimate the age of a biological sample found at a crime scene. RNA sequencing of transcripts from various tissue types shows that degradation occurs faster at the 5’ end than the 3’ end. This discovery led to the development of the 5’-3’ assay, which quantifies and compares each end of a transcript in a single reaction to estimate sample age. This assay has been validated on dried bloodstains, however why the 5’ end of the transcript degrades faster than the 3’ end remains unknown. As this phenomenon is being observed in dried samples, we hypothesize that chemical hydrolysis reactions are responsible for breaking the RNA molecule and thus chemical modifications of the 5’ or 3’ ends of the RNA molecule may affect the degradation rate. A literature exists that suggests that methylation of RNA molecules can alter the kinetics of RNA degradation through affecting transcript stability and also possibly dependent on the RNA binding proteins (RBPs) present. We aim to investigate the methylation patterns of transcripts in dried bloodstains using RNA enrichment and liquid chromatography tandem mass spectrometry (LC-MS/MS). We developed a novel RNA enrichment technique that utilizes 120bp DNA probes designed to hybridize to the 5’ or 3’ ends of a transcript for selected target enrichment. The enriched product will then be hydrolyzed into nucleotides for analysis via LC-MS/MS. Preliminary results show that this approach can enrich for our selected target over 150,000-fold. We anticipate observing differences in RNA methylation patterns between the 5’ and 3’ ends of our selected transcripts, potentially explaining the differential degradation rates of the 5’ and 3’ ends of RNA molecules. | |
| dc.format | application/pdf | |
| dc.language | en_US | |
| dc.publisher | Oklahoma State University Center for Health Sciences | |
| dc.rights | The author(s) retain the copyright or have the right to deposit the item giving the Oklahoma State University Library a limited, non-exclusive right to share this material in its institutional repository. Contact Digital Resources and Discovery Services at lib-dls@okstate.edu or 405-744-9161 for the permission policy on the use, reproduction or distribution of this material. | |
| dc.title | Detection of RNA methylation patterns of forensically relevant transcripts in dried bloodstains | |
| osu.filename | ouhd_Lohr_detectionofRNAmethylation_2022.pdf | |
| dc.type.genre | Presentation | |
| dc.type.material | Text | |
| dc.subject.keywords | RNA degradation | |
| dc.subject.keywords | RNA methylation | |
| dc.subject.keywords | qPCR | |